This website uses cookies so that we can provide you with the best user experience possible. Cookie information is stored in your browser and performs functions such as recognising you when you return to our website and helping our team to understand which sections of the website you find most interesting and useful.
Hack Your Health: An Evening with Anjali Nayar and Dr. Sean Gibbons
at the Institute for Systems Biology
Hack Your Health: An Evening with Anjali Nayar and Dr. Sean Gibbons
Netflix’s “Hack Your Health: The Secrets of Your Gut” is a documentary that merged gut microbiome experts, four individuals – including a well-known hot dog eating champion – facing personal battles with gastrointestinal health, and a unique, effective visual method of “showing” the gut microbiome in action.

‘1 in 1,000:’ Dr. Sean Gibbons Named Highly Cited Researcher for 2023
ISB Associate Professor Dr. Sean Gibbons was named a Highly Cited Researcher for 2023. It is the second consecutive year Gibbons has earned the distinction. The Highly Cited Research list is generated annually by Clarivate, which says: “Of the world’s population of scientists and social scientists, Highly Cited Researchers are 1 in 1,000.”

ISB and Seattle Science Foundation Partner to Create Video Series
What are multi-omics? Why does our microbiome matter? What’s the difference between genetics and genomics? What is a digital twin? ISB and Seattle Science Foundation have partnered to create videos answering questions like these and more, showcasing ISB scientists and their work.
Bugs vs. Drugs: How Our Microbiomes Can Explain Our Response to Statins
ISB Assistant Professor Dr. Sean Gibbons talked about the science behind statins in our most recent Research Roundtable virtual presentation. His talk was titled “Bugs vs. Drugs: How Our Unique Gut Microbiomes Shape Our Personalized Responses to Statins.”
Dr. Jack Gilbert on the State of the Microbiome Field
In the final ISB-Town Hall Seattle Science Series of 2021, ISB Assistant Professor Dr. Sean Gibbons sat down with UCSD Professor Dr. Jack Gilbert, and the two microbiome experts discussed past research, exciting science happening today, promising products and therapies on the horizon, and much more.
Personalized Nutrition and Your Gut Microbiome
In ISB’s first-ever Research Roundtable event, Assistant Professor Dr. Sean Gibbons delivered a presentation titled “Gut-Check: Personalized Nutrition and Your Microbiome.” His talk covered a lot of ground, including recently published research showing how the health of our microbiomes can predict longevity, and how we can build and maintain a healthy gut microbiome.
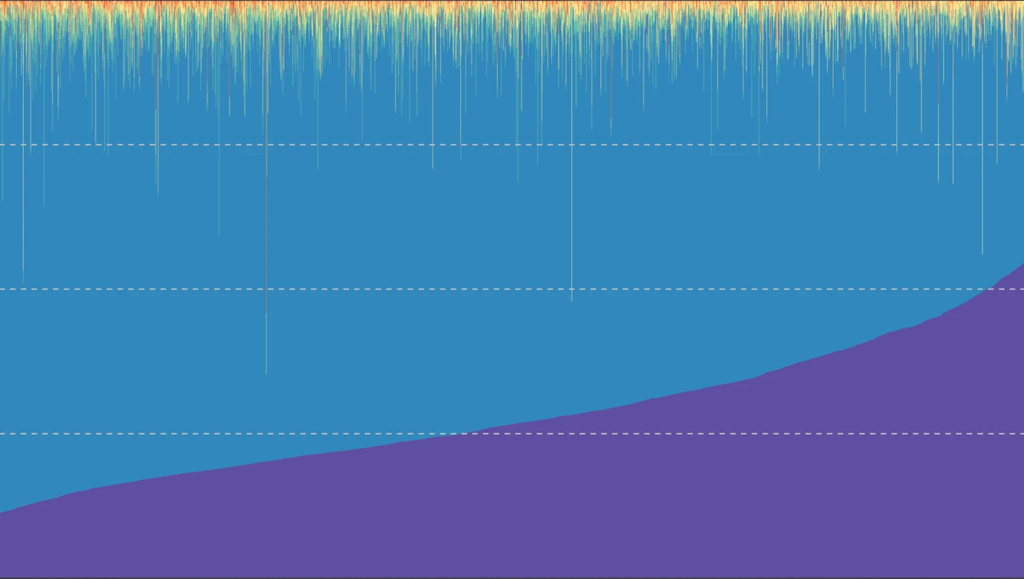
Variations in the Microbiome Associated with Health, Disease
ISB researchers examined the associations between the gut microbiomes of about 3,400 people and roughly 150 host characteristics. The team looked at diet, medication use, clinical blood markers, and other lifestyle and clinical factors, and found evidence that variations of the gut microbiome are associated with health and disease.
Answering Nature’s Call: How Scientists Are Mining Sewage To Track Population Health
Everybody pees and poops. What if there was a way to use the byproducts of our everyday bodily functions to understand the general health of a population? That is exactly what MIT’s Dr. Eric Alm is pursuing. In an ISB-Town Hall Seattle live stream, Alm discussed the promise of this novel form of public health tracking.

Harnessing our inner ecology to track and treat disease
ISB’s virtual course and symposium focusing on the microbiome and its future role in precision medicine will take place on October 15 and 16. The event’s website went live earlier this week. The virtual course will be taught by Sean Gibbons, Christian Diener, Tomasz Wilmanski, Noa Rappaport, Alex Carr, Priyanka Baloni and Nathan Price. Symposium speakers are Jason Papin (University of Virginia), Ines Thiele (National University of Ireland, Galway), Thomas…

ISB Microbiome Researcher Dr. Sean Gibbons Featured in TIME Article
Freaked out about a “germy” bathroom? You don’t need to be. ISB Assistant Professor and microbiome researcher Dr. Sean Gibbons was featured prominently in an article, headlined “The Germiest Place in your Bathroom Isn’t Your Toilet,” published online by TIME.

All About the Human Microbiome
The human microbiome is a relatively new area of research, and there are numerous questions surrounding it. What is the human microbiome? Can we change it? Does it make us sick? Keep us well? ISB Assistant Professor and microbiome researcher Dr. Sean Gibbons answers these questions — and many more.

ISB's Dr. Sean Gibbons on the importance of the human microbiome
“This new organ that we’re coming to recognize as the microbiome is part and parcel to the functionality of the whole system, and if it breaks down, if it starts to fall apart, we start to get sick,” said Dr. Sean Gibbons, ISB’s newest faculty member, in a WGBH Forum Network presentation.