This website uses cookies so that we can provide you with the best user experience possible. Cookie information is stored in your browser and performs functions such as recognising you when you return to our website and helping our team to understand which sections of the website you find most interesting and useful.
Timing is Everything: ISB Study Finds Link Between Bowel Movement Frequency and Overall Health
at the Institute for Systems Biology

Timing is Everything: ISB Study Finds Link Between Bowel Movement Frequency and Overall Health
Everybody poops, but not every day. An ISB-led research team examined the clinical, lifestyle, and multi-omic data of more than 1,400 healthy adults. How often people poop, they found, can have a large influence on one’s physiology and health.
A New Path Toward Microbiome-Informed Precision Nutrition
ISB researchers have developed a novel way to simulate personalized, microbiome-mediated responses to diet. They use a microbial community-scale metabolic modeling (MCMM) approach to predict individual-specific short-chain fatty acid production rates in response to different dietary, prebiotic, and probiotic inputs.

‘1 in 1,000:’ Dr. Sean Gibbons Named Highly Cited Researcher for 2023
ISB Associate Professor Dr. Sean Gibbons was named a Highly Cited Researcher for 2023. It is the second consecutive year Gibbons has earned the distinction. The Highly Cited Research list is generated annually by Clarivate, which says: “Of the world’s population of scientists and social scientists, Highly Cited Researchers are 1 in 1,000.”
Gut Microbiome Composition Predictive of Patient Response to Statins
New ISB research shows that different patient responses to statins can be explained by the variation in the human microbiome. The findings were published in the journal Med, and suggest that microbiome monitoring could be used to help optimize personalized statin treatments.
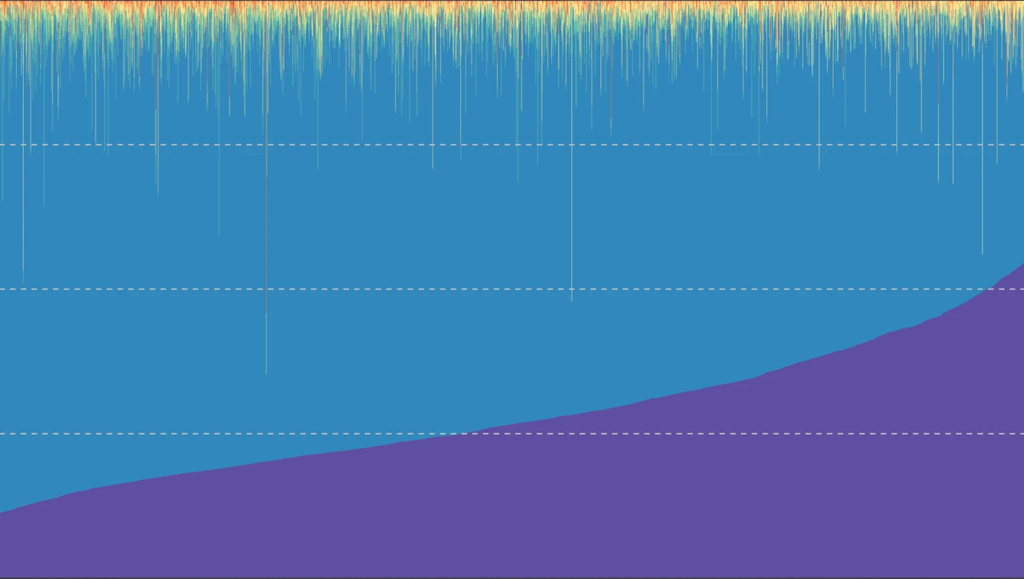
Variations in the Microbiome Associated with Health, Disease
ISB researchers examined the associations between the gut microbiomes of about 3,400 people and roughly 150 host characteristics. The team looked at diet, medication use, clinical blood markers, and other lifestyle and clinical factors, and found evidence that variations of the gut microbiome are associated with health and disease.